一、安装与启用
Click on the green "Clone or download" button on the top right corner of this page to obtain the web URL. Download this module by running git clone <url>on command line.</url>
将clone的文件放在 /sites/all/modules下面,然后去启用模块
Tripal Expression(在TRIPAL EXTENSIONS下面)的依赖模块如下:Requires: Tripal (enabled), Views (enabled), Chaos tools (enabled), Path (enabled), Search (enabled), PHP filter (enabled), Entity API (enabled), Redirect (enabled), Tripal Chado (enabled), Date (enabled), Date API (enabled), Image (enabled), File (enabled), Field (enabled), Field SQL storage (enabled), Link (enabled), Tripal Chado Views (enabled), Tripal Biomaterials (disabled), jQuery Update (enabled)
所以还要下载安装开启一个Tripal Biomaterials的模块,可以去容器直接复制拖过来91容器/home/shicy/web/modules/tripal_analysis_expression-master,同时在网站上打开模块Tripal Biomaterials,Tripal Expression两个模块
```
[shicy@localhost tripal_analysis_expression-master]$ docker cp /home/shicy/web/modules/tripal_analysis_expression-master 2175945e3164:/var/www/html/sites/all/modules/
```
二、前期文件准备(在92的wzz_python容器中进行)
需要香榧的biosample数据xml文件和表达量csv文件以及转录本fasta文件
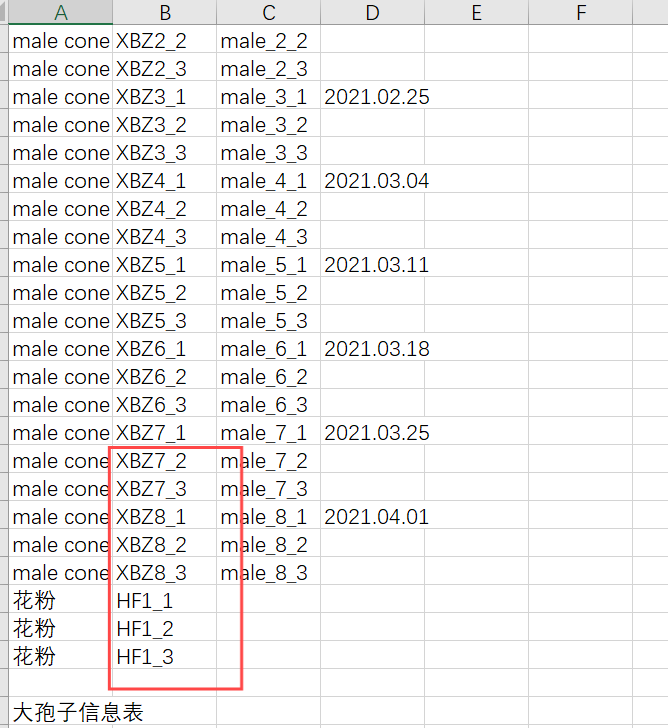
1、先将csv文件修改成如下格式。将对应原始数据放在第二列,命名放在第三列,描述放在第一列。
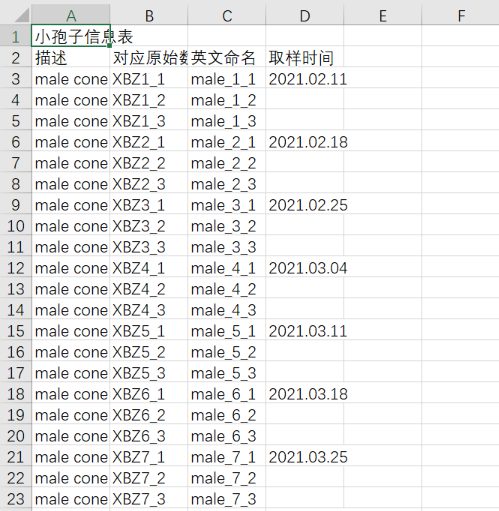
2、biosample文件需要自己通过csv文件生成。写一个python代码,generate_biosample_xml.py。这个代码没有办法完全自动生成,后续需要手动改。
```
root@aee56b84d678:/home/scy/ba_expression_transcripts/expression_transcript_ba# cat generate_biosample_xml.py
import csv
import xml.etree.ElementTree as ET
# 创建根元素
biosample_set = ET.Element('BioSampleSet')
# 读取CSV文件并生成BioSample元素
with open('FPKM-信息表.csv', newline='', encoding='iso-8859-1') as csvfile:
reader = csv.reader(csvfile)
next(reader) # 跳过标题行
for row in reader:
if not row[0]: # 如果第一列为空,跳过
continue
# 创建BioSample元素
biosample = ET.SubElement(biosample_set, 'BioSample', access="public", accession=row[1])
# 创建Ids元素
ids = ET.SubElement(biosample, 'Ids')
# 将CSV文件中“原始数据文件名”列的值映射到Id元素的文本内容
ET.SubElement(ids, 'Id', db="BioSample", is_primary="1").text = row[1] # “原始数据文件名”是第二列
ET.SubElement(ids, 'Id', db_label="Sample name").text = row[2] # “样本名称”是第一列
# 创建Description元素
description = ET.SubElement(biosample, 'Description')
ET.SubElement(description, 'Title').text = row[0] # “原始数据文件名”是第二列
organism = ET.SubElement(description, 'Organism', taxonomy_name="Torreya grandis")
ET.SubElement(organism, 'OrganismName').text = "Torreya grandis"
comment = ET.SubElement(description, 'Comment')
# 创建Owner元素
owner = ET.SubElement(biosample, 'Owner')
ET.SubElement(owner, 'Name', url="https://www.zafu.edu.cn/").text = "Zhejiang A&F University"
# 创建Models元素
models = ET.SubElement(biosample, 'Models')
ET.SubElement(models, 'Model').text = "Plant"
# 创建Package元素
ET.SubElement(biosample, 'Package', display_name="Plant; version 1.0").text = "Plant.1.0"
# 将生成的XML写入文件
tree = ET.ElementTree(biosample_set)
tree.write('biosample_result.xml', encoding='utf-8', xml_declaration=True)
root@aee56b84d678:/home/scy/ba_expression_transcripts/expression_transcript_ba# python generate_biosample_xml.py
root@aee56b84d678:/home/scy/ba_expression_transcripts/expression_transcript_ba# ls
FPKM-信息表.csv gene_count_matrix_other.csv gene_tpm_matrix.csv
biosample_result.xml gene_fpkm_matrix.csv gene_tpm_matrix_fruit.csv
gene_count_matrix.csv gene_fpkm_matrix_fruit.csv gene_tpm_matrix_other.csv
gene_count_matrix_fruit.csv gene_fpkm_matrix_other.csv generate_biosample_xml.py
```
注意:这里要注意的是,csv文件的名字和表达量的文件名字需要一致。
3、转录本fasta文件:061b51fa0bb3:/home/scy/Torreya_grandis/torreya_noptg/transcripts.fasta
转录本fasta文件可以用gffread命令通过基因组fasta文件和gff3注释文件提取,提取命令:
```
root@061b51fa0bb3:/home/scy/Torreya_grandis/torreya_noptg# gffread -w tanscripts.fasta -g Tgra_noptg.fa Tgra_noptg.gff
No fasta index found for Tgra_noptg.fa. Rebuilding, please wait..
Fasta index rebuilt.
root@061b51fa0bb3:/home/scy/Torreya_grandis/torreya_noptg# ls -Allthr
total 18G
-rw-r--r-- 1 1010 1011 18G Dec 18 13:04 Tgra_noptg.fa
-rw-r--r-- 1 1010 1011 713 Dec 18 13:22 noptg_gff.py
-rw-r--r-- 1 1010 1011 30M Dec 18 13:22 Tgra_noptg.gff
-rw-r--r-- 1 1010 1011 626 Dec 18 13:30 noptg_pep.py
-rw-r--r-- 1 1010 1011 12M Dec 18 13:30 Tgra_noptg_pep.fa
-rw-r--r-- 1 root root 537 Dec 30 2024 Tgra_noptg.fa.fai
-rw-r--r-- 1 root root 35M Dec 30 2024 transcripts.fasta
```
4、原始csv文件存储位置:/data2/zhenzixu/rnaseq。后续处理完生成biosample的数据在92的/data2/shicy/raw/transcript_ba/expression_transcript_ba中
三、上传数据(91测试容器torreya_ba,端口5588)
1、上传香榧的biosample数据数据
这里有可能报错,要把/var/www/html/sites/default/files/tripal/users文件和users文件下面的1文件的权限都改成apache.apache
[root@9c513d1535e3 tripal]# ls -Allthr
total 0
drwxrwxrwx 3 apache apache 23 Mar 26 12:37 users
drwxr-xr-x 2 root root 10 Mar 26 12:37 tripal_analysis_expression
drwxr-xr-x 2 root root 10 Mar 26 12:37 tripal_analysis_expression_download
drwxr-xr-x 2 root root 10 Mar 26 12:37 chado_search
[root@9c513d1535e3 users]# ls -Allthr
total 0
drwxr-xr-x 3 apache apache 94 Mar 26 12:55 1(1)在Tripal -> DataLoader -> Chado Biological Sample (Biomaterial) Loader中上传数据,选择右下角的按钮执行下一步直到submit。数据在/data2/shicy/raw/transcript_ba/expression_transcript_ba/biosample_result.xml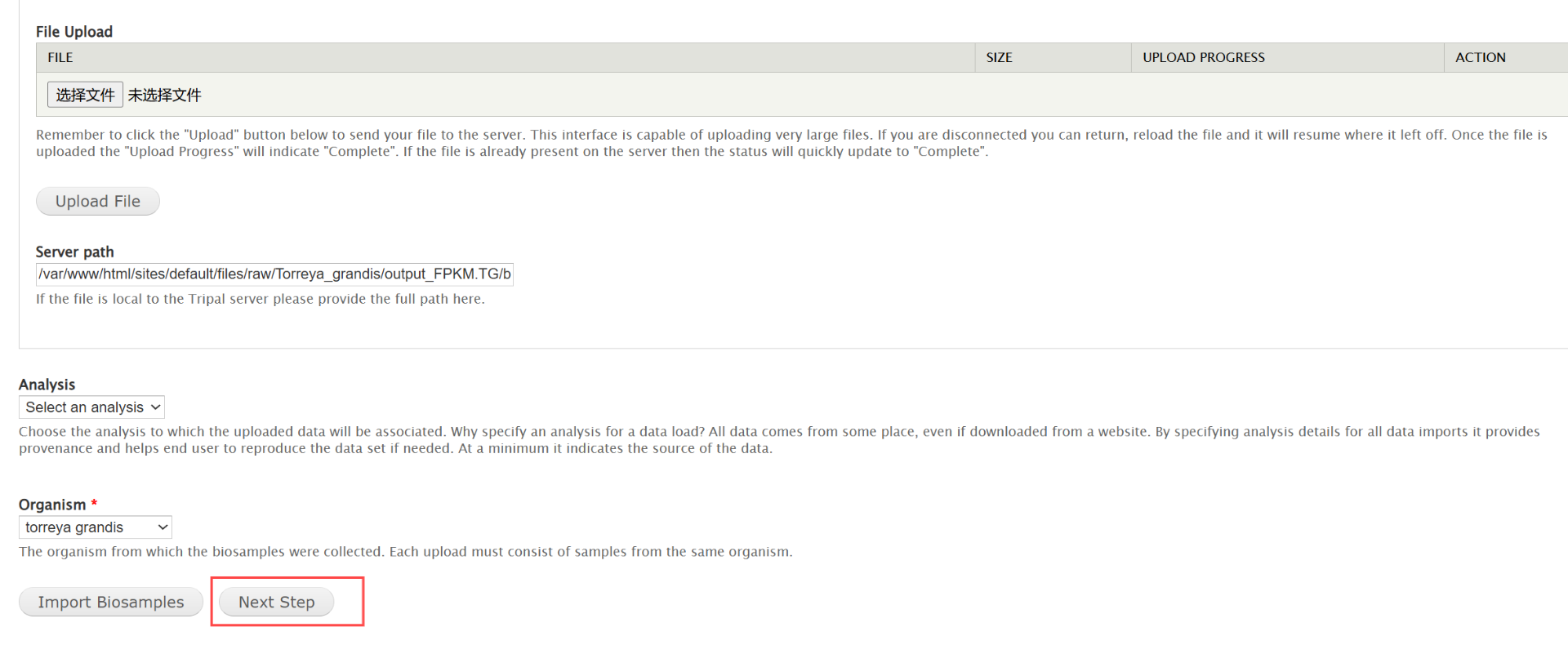
在服务器中运行drush命令
[root@9c513d1535e3 users]# drush trp-run-jobs --username=admin --root=/var/www/html
2025-03-26 13:18:44
Tripal Job Launcher
Running as user 'admin'
-------------------
2025-03-26 13:18:44: There are 4 jobs queued.
2025-03-26 13:18:44: Job ID 201.
2025-03-26 13:18:44: Calling: tripal_tripal_cron_notification()
2025-03-26 13:18:47: Job ID 202.
2025-03-26 13:18:47: Calling: tripal_expire_collections()
2025-03-26 13:18:47: Job ID 203.
2025-03-26 13:18:47: Calling: tripal_expire_files()
2025-03-26 13:18:47: Job ID 204.
2025-03-26 13:18:47: Calling: tripal_run_importer(168)
Running 'Chado Biological Sample (Biomaterial) Loader' importer
NOTE: Loading of file is performed using a database transaction.
If it fails or is terminated prematurely then all insertions and
updates are rolled back and will not be found in the database
Opening temporary cache file: /tmp/TripalBiomaterialImport_3v36CV
Step 1 of 10: Caching input file...
Found 130 samples
Step 2 of 10: Find existing samples...
Step 3 of 10: Clear attributes of existing samples...
Step 3 of 10: Insert samples...
Step 5 of 10: Find sample IDs...
Step 6 of 10: Insert sample properties...
Step 7 of 10: Get cross references...
Step 8 of 10: Insert new cross references...
Step 9 of 10: Get new cross references IDs...
Step 10 of 10: Insert sample cross references...
Removing temporary cache file: /tmp/TripalBiomaterialImport_3v36CV
Done.
Remapping Chado Controlled vocabularies to Tripal Terms...
Done.再在网页上的Home » Administration » Content » Tripal Content » Publish Tripal Content中选择Biological Sample,publish之后再去网页上drush一下
[root@9c513d1535e3 users]# drush trp-run-jobs --username=admin --root=/var/www/html
2025-03-26 13:19:19
Tripal Job Launcher
Running as user 'admin'
-------------------
2025-03-26 13:19:19: There are 1 jobs queued.
2025-03-26 13:19:19: Job ID 205.
2025-03-26 13:19:19: Calling: chado_publish_records(Array)
INFO (PUBLISH_RECORDS): There are 130 records to publish.
There are 130 records to publish.
INFO (PUBLISH_RECORDS): Successfully published 130 Biological Sample record(s).
Successfully published 130 Biological Sample record(s).
2、上传fasta文件
在Home » Administration » Tripal » Data Loaders » Chado FASTA Loader中上传fasta文件,注意Sequence Type选择mrna
[root@2175945e3164 transcript_ba]# drush trp-run-jobs --username=admin --root=/var/www/html
2025-03-26 04:48:38
Tripal Job Launcher
Running as user 'admin'
-------------------
2025-03-26 04:48:38: There are 1 jobs queued.
2025-03-26 04:48:38: Job ID 206.
2025-03-26 04:48:38: Calling: tripal_run_importer(169)
Running 'Chado FASTA Loader' importer
NOTE: Loading of file is performed using a database transaction.
If it fails or is terminated prematurely then all insertions and
updates are rolled back and will not be found in the database
Step 1: Finding sequences...
Step 2: Importing sequences...
Found 57074 sequence(s).
Percent complete: 100.00 %. Memory: 62,143,344 bytes.
Done.
Remapping Chado Controlled vocabularies to Tripal Terms...
Done.
在网页上的Home » Administration » Content » Tripal Content » Publish Tripal Content选择mRNA,publish之后在服务器drush一下命令。因为之前上传过mrna数据
[root@2175945e3164 transcript_ba]# drush trp-run-jobs --username=admin --root=/var/www/html
2025-03-26 05:30:28
Tripal Job Launcher
Running as user 'admin'
-------------------
2025-03-26 05:30:28: There are 1 jobs queued.
2025-03-26 05:30:28: Job ID 207.
2025-03-26 05:30:28: Calling: chado_publish_records(Array)
INFO (PUBLISH_RECORDS): There are 0 records to publish.
There are 0 records to publish.
INFO (PUBLISH_RECORDS): Successfully published 0 mRNA record(s).
Successfully published 0 mRNA record(s).
3、上传csv文件
在网页上Home » Administration » Tripal » Data Loaders » Chado Expression Data Loader中上传csv文件。再去服务器中drush一下命令即上传成功。
**注意**:这里上传文件容易报错,因为用户数据目录没有赋予写入权限,于是去sites目录下面找,原来tripal的用户数据目录是在sites/default/files/tripal目录中,于是给它开放写入权限,即可上传成功数据
上传csv文件页面时需要的配置
- **分析** - 要与表达式数据关联的分析。
- **Organism (required)** - 生物体。
- **Sequence Type** - mRNA
- **源文件类型** - Matrix Format
- **名称匹配类型** - uniquename
- **Data Delimiter** - comma
- **文件类型后缀** - 要加载的文件的后缀。这用于在同一目录中提交多个列格式文件。矩阵文件不需要后缀。
- **数据开头的正则表达式** - 如果表达式文件具有标头,则使用此字段捕获表达式数据开头之前出现的行。此行文本和此行之前的任何文本都将被忽略。
- **数据末尾的正则表达式** - 如果表达式文件具有页脚,则使用此字段捕获表达式数据末尾之后出现的行。此行文本和后面的所有文本都将被忽略。
- **阵列设计** - 这仅适用于微阵列表达数据。对于不使用阵列的实验(即下一代测序),这可能会留空。
- **单位** - 与加载值关联的单位,例如 FPKM。您也可以使用**定量单位**管理页面更新实验单位。
把所有csv文件上传成功。
4、在网页上Home » Administration » Structure » Blocks打开相应模块tripal_analysis_expression features form for heatmap和tripal_analysis_expression heatmap display
5、Example修改的位置在Tripal ->Extensions -> Expression Analysis -> Expression Heatmap Search Settings,上面填写的csv文件第一列的id号。我这里填写的是mrna_id
复现成功